Our variant calling pipeline is adapted for germline or somatic variants from panels, exomes or genomes. To assure the best results, the samples data firstly pass through a rigorous quality control, and the final results are extensively annotated using more than 30 different databases (population frequency, function impact, conservation, COSMIC, ClinVar etc), useful for a better data interpretation.
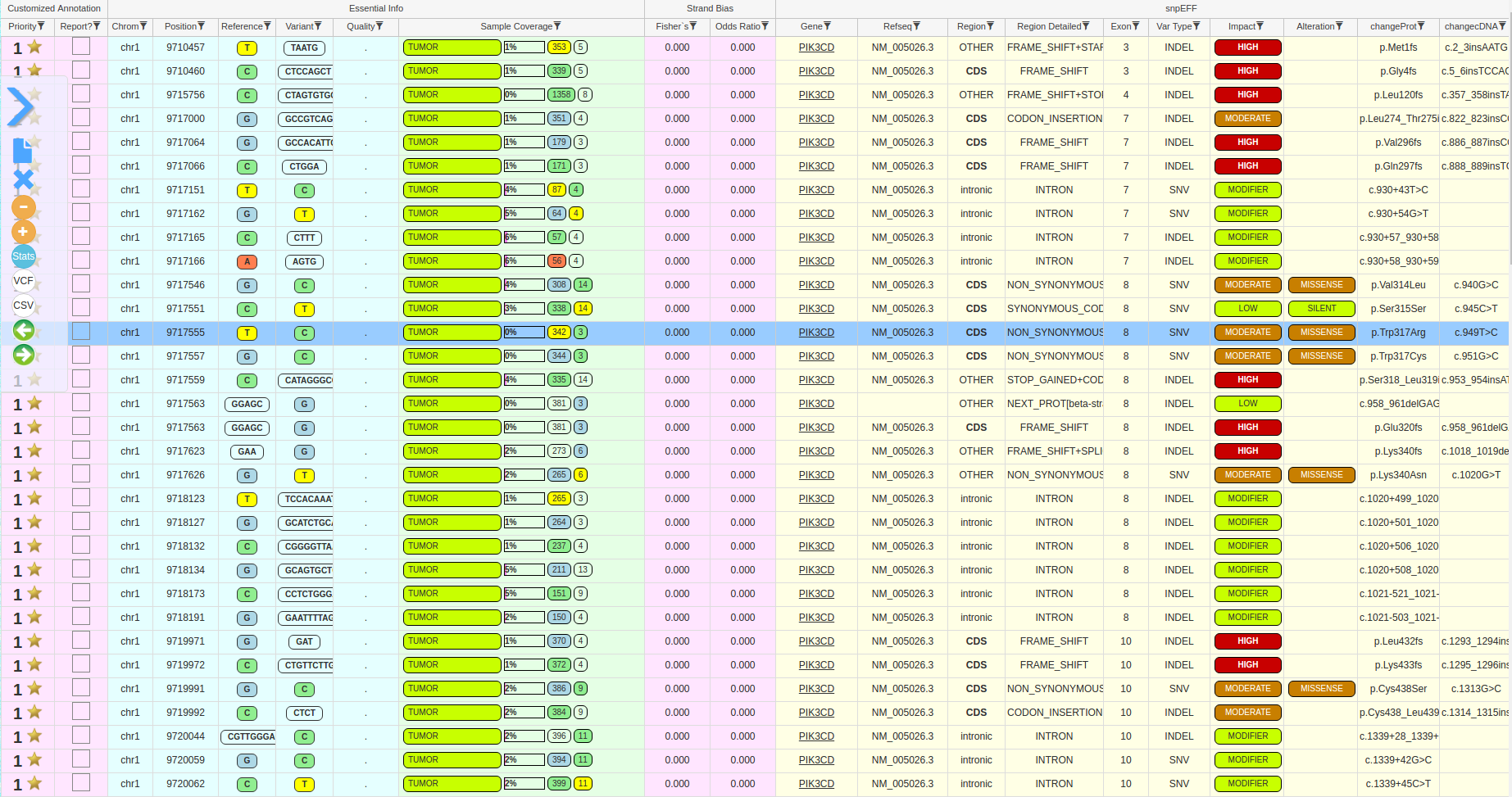
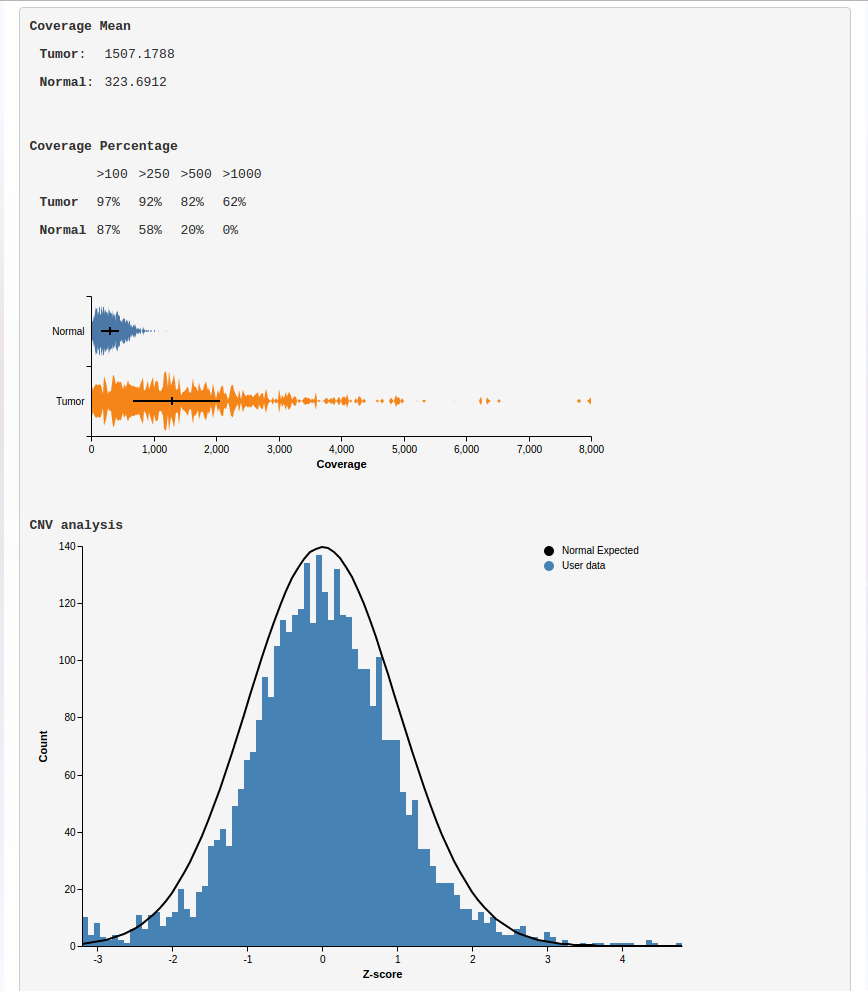
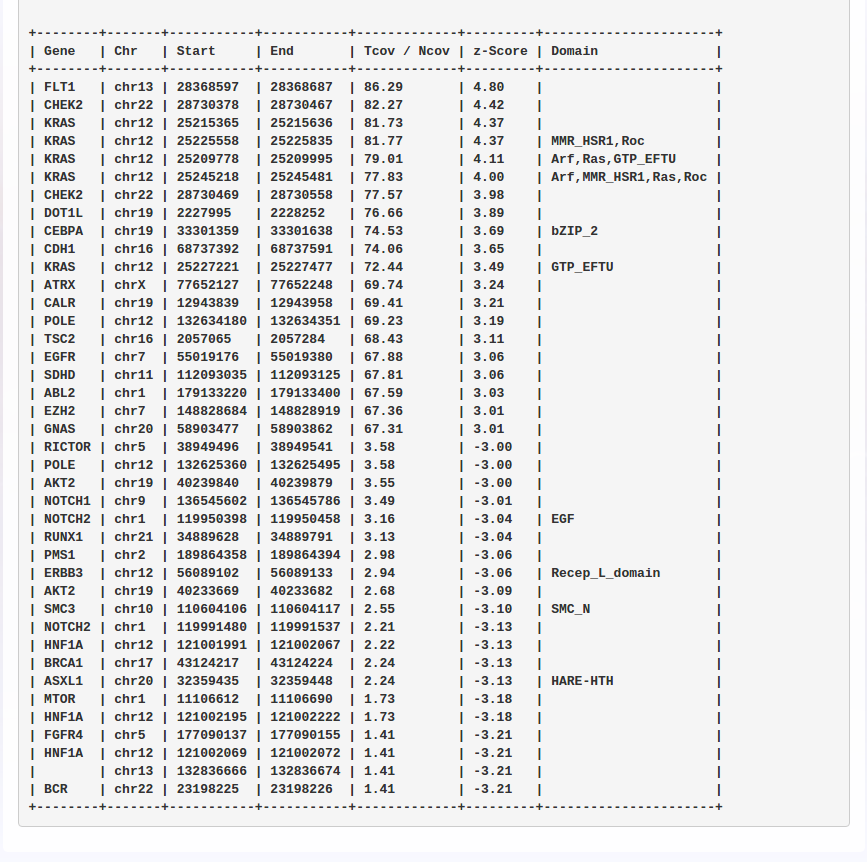
The results are provided through a variant viewer (ViVa – Viewer of Variants), which is an advanced tool that allows the user to manage the variants and execute complex filterings and/or sample comparisons. A small report is also provided to the user, showing general quality metrics regarding the analyzed samples.